Description
INTRODUCTION: The recombinant respiratory syncytial viruses (RSV) introduced by ViraTree including RSV-GFP1 (Product #R121), RSV-RFP1 (#R131), and RSV-GFP1-Luc2 (#R141), have become very popular among wide range of investigators, however, these first generation viruses have limitations. One is that the location of the reporter genes (GFP, RFP, and/or Luc) in the RSV genome (in front of NS1 gene) is not optimal for virus propagation. Secondly, the reporter fluorescent proteins in these viruses are of earlier versions, and brighter and more stable counterparts that are optimized for mammalian cell expression have become available. Therefore the second generation recombinant RSVs were created by 1) replacing the original GFP or RFP reporters with the improved versions with enhanced fluorescence and expression profiles, and 2) moving the reporter gene from the first position (in front of NS1) downstream to between the P and M genes. The position between the P and M genes was chosen because it places the foreign insert well downstream of the NS1 and NS2 genes, avoiding direct effects on the expression of NS1 and NS2. To construct a transcriptional cassette for the reporter gene, the G gene end signal was chosen because it is among the most efficient of these signals and would have the minimum impact on the viral transcriptional program; the gene start signals are more highly conserved, and that of N is typical and was used. The second generation recombinant RSVs include RSV-GFP5 (Product #R125), RSV-DsRed5 (#R135), and RSV-FarRed5 (#R155). RSV-GFP5 expresses enhanced GFP (EGFP, Clontech) which is both brighter and more stable comparing to the original GFP. Together these novel viruses are suitable for multicolor applications and/or whole body imaging.
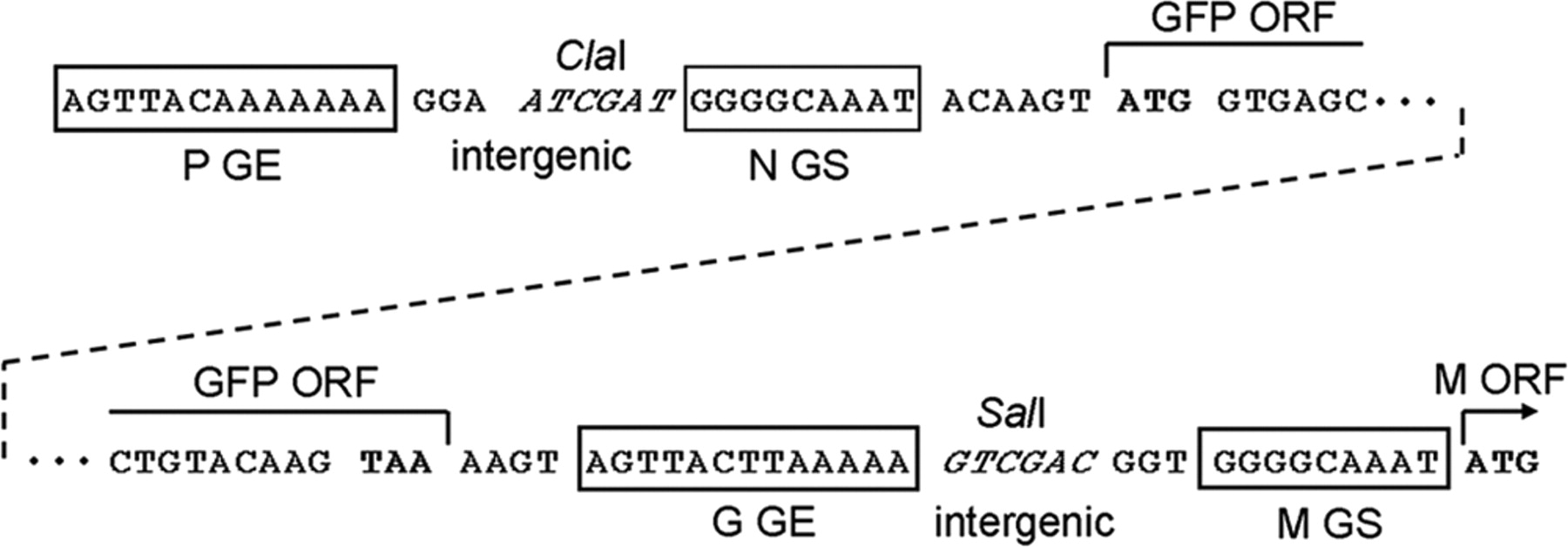
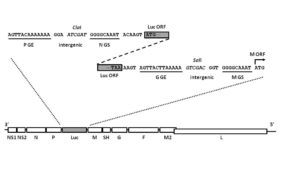
DESCRIPTION: Wild-type parent based on RSV strain A2 were modified to express enhanced GFP from a gene cassette placed between the P and M genes. The open reading frame (ORF) for enhanced green fluorescent protein (EGFP, Clontech) was PCR amplified to add a ClaI site and an RSV gene start signal (from the N gene) upstream of the ORF and an RSV gene end signal (from the G gene) and SalI site downstream of the ORF. The ClaI-SalI fragment was further cloned through multiple steps between the P and M genes in the full-length RSV cDNA plasmid, in which silent mutations were introduced into the last three codons of the SH protein and the stop codon and 112 nucleotides of noncoding region of the SH gene downstream the ORF were removed in order to stabilize the plasmid for propagation in bacteria. Recombinant RSV-GFP5 was rescued by transfecting T7 polymerase-expressing BSR T7/5 cells with the full-length antigenome plasmid and plasmids expressing N, P, L, and M2-1 genes. The recovered RSV-GFP5 was further propagated in HEp-2 cells, the expression of GFP by infected cells was confirmed by fluorescence microscopy, and the sequence of the virus was confirmed in its entirety. RSV-GFP5 was found to replicate as efficiently as its biological parental strain in cell cultures.
| Parental Strain: | Strain A2 |
| Construction: | GFP was inserted between P and M genes as the 5th gene. |
| Passage History: | The isolate was propagated in HEp-2 cells (ATCC CCL-23). |
| Infectivity: | Titer > 6.5 log10 TCID50 per mL. |
| Volume/Storage: | 2 x 1.2 mL per cryovial. Store at –80ºC. |
| Quality Testing: | No bacteria, fungus, or mycoplasma detected. Endotoxin <10 EU/mL. |
| Availability: | Bulk quantity and custom orders are available. Contact info@viratree.com. |
Product Sheet: Product_R125
Certificate of Analysis: CoA_Lot-2507
Material Safety Data Sheet (MSDS): MSDS_RSV